Viral lineages were classified by using the Pangolin ( 26) software tool ( ), nextclade ( ), and standard phylogenetic analysis using complete reference genomes. 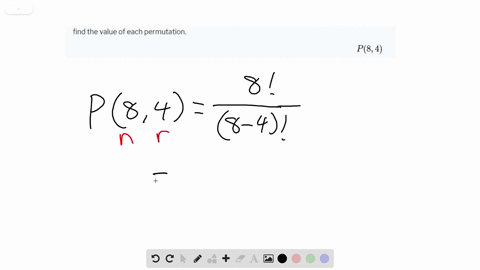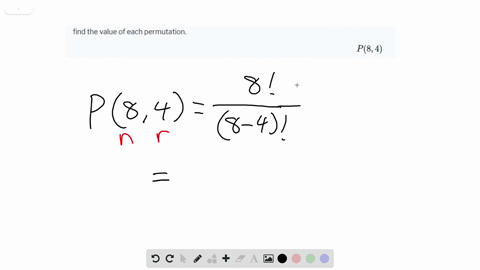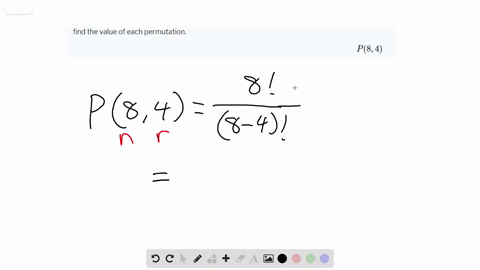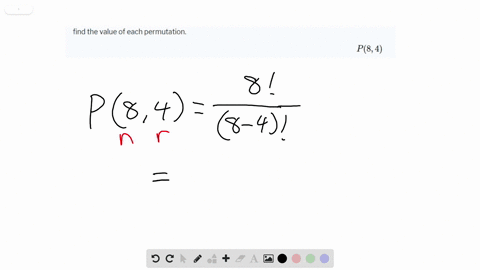
S1 to S4), together with other available and published genomes from Brazil for context. Because partial genome sequences can provide useful epidemiological information, particularly regarding virus genetic diversity and lineage composition ( 25), we harnessed information from partial ( n = 41 viral sequences, 25 to 75% genome coverage), as well as near-complete ( n = 95 viral sequences, 75 to 95%) and complete ( n = 48 viral sequences, ≥95%) sequences from Manaus (figs.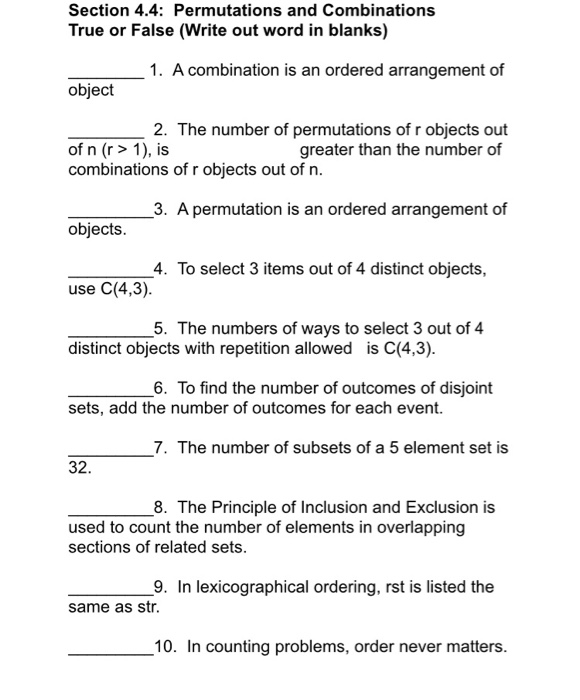 
We sequenced SARS-CoV-2 genomes from 184 samples from patients seeking COVID-19 testing in two diagnostic laboratories in Manaus between November and December 2020, using the ARTIC V3 multiplexed amplicon scheme ( 24) and the MinION sequencing platform. Before this, only seven SARS-CoV-2 genome sequences from Amazonas were publicly available (SARS-CoV-2 was first detected in Manaus on 13 March 2020) ( 19, 23). 1A), we focused ongoing SARS-CoV-2 genomic surveillance ( 2, 18– 22) on recently collected samples from the city (supplementary materials, materials and methods, and table S2). Enhanced global genomic surveillance of variants of concern, which may exhibit increased transmissibility and/or immune evasion, is critical to accelerate pandemic responsiveness.Īfter a rapid increase in hospitalizations in Manaus caused by severe acute respiratory infection (SARI) in December 2020 ( Fig. Using a two-category dynamical model that integrates genomic and mortality data, we estimate that P.1 may be 1.7- to 2.4-fold more transmissible and that previous (non-P.1) infection provides 54 to 79% of the protection against infection with P.1 that it provides against non-P.1 lineages. Molecular clock analysis shows that P.1 emergence occurred around mid-November 2020 and was preceded by a period of faster molecular evolution. Lineage P.1 acquired 17 mutations, including a trio in the spike protein (K417T, E484K, and N501Y) associated with increased binding to the human ACE2 (angiotensin-converting enzyme 2) receptor. Genome sequencing of viruses sampled in Manaus between November 2020 and January 2021 revealed the emergence and circulation of a novel SARS-CoV-2 variant of concern. Coupled with the emergence of P.1, disease spread was accelerated by stark local inequalities and political upheaval, which compromised a prompt federal response.Ĭases of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection in Manaus, Brazil, resurged in late 2020 despite previously high levels of infection. 
After a period of accelerated evolution, this variant emerged in Brazil during November 2020. tracked the evolution of a new, more aggressive lineage called P.1, which has 17 mutations, including three (K417T, E484K, and N501Y) in the spike protein. In Manaus, transmission reached unprecedented levels after a momentary respite in mid-2020. SARS-CoV-2 circulated undetected in Brazil for more than a month as it spread north from Sã o Paulo. Clusters of deaths before cases became apparent indicated unmitigated spread. analyzed the pattern of spread of COVID-19 cases and deaths in the country from February to October 2020. Using daily data from state health offices, Castro et al. Sabino +69 authors +67 authors +62 authors fewer Authors Info & Affiliationsĭespite an extensive network of primary care availability, Brazil has suffered profoundly during the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic. Pybus, Seth Flaxman, Samir Bhatt, and Ester C. Loman, Philippe Lemey, Andrew Rambaut, Nelson A. Pond, Chieh-Hsi Wu, Oliver Ratmann, Neil M. Hay, Mélodie Monod, Xenia Miscouridou, Helen Coupland, Raphael Sonabend, Michaela Vollmer, Axel Gandy, Carlos A. Camilo, Henrique Hoeltgebaum, William M. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
